What is it about?
This commentary summarises recent progress in linking the DNA sequence of bacteria which cause infections to the antibiotics they're resistant to can help us design better diagnostics to make better decisions about which antibiotics to prescribe to which patients. We outline some studies which have used statistical association testing to link variation in DNA to resistance, but also highlight some recent applications of machine learning which may allow us to identify resistance mechanisms missed by traditional tests.
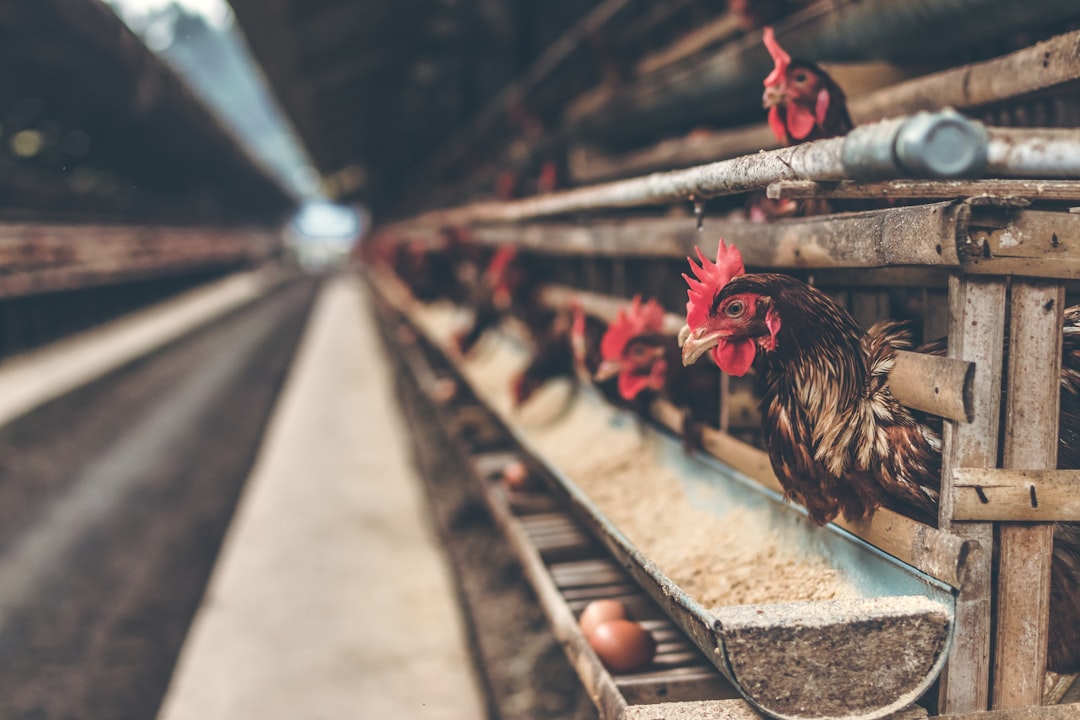
Featured Image
Photo by Mark Fletcher-Brown on Unsplash
Why is it important?
Currently, lab-based technologies are being used to identify antibiotic resistance, but over time we will see a movement to using whole-genome sequencing (WGS), because of the additional benefits in allowing us to identify transmission of infections, disease outbreaks, and the emergence and spread of new resistance mechanisms. Progress like that identified in these recent studies is key to driving the reliability of WGS-based tests and encouraging uptake of sequencing in public health labs.
Perspectives
This commentary highlights the great progress we're making in identifying dangerous strains of bacteria with DNA sequencing. We also note the surprising observation that a machine learning algorithm trained with all elements of the genome except those known to cause resistance to antibiotics can predict resistance just as effectively as one trained with only elements known to cause resistance.
Dr Nicole E Wheeler
Wellcome Sanger Institute
Read the Original
This page is a summary of: Breaking the code of antibiotic resistance, Nature Reviews Microbiology, March 2018, Springer Science + Business Media,
DOI: 10.1038/nrmicro.2018.33.
You can read the full text:
Contributors
The following have contributed to this page










